SIM - Image Gallery
The University of Arizona added a Structured Illumination Microscope (Zeiss Elyra S.1) to it's microscopy toolbox in 2016. SIM uses patterned illumination, projected from multiple angles, to achieve resolution that is 2X better than that of regular confocal or deconvolution microscopes.
The resolution improvement from SIM does have some caveats. Careful attention to sample preparation details is crucial. The good news is these same details will improve the images acquired on any fluorescence microscope (widefield, deconvolution, confocal). There are limits (approx 15um) to how deep into a sample we can SIM image. In our experience, some samples work beautifully (typically those with bright, small, discrete structures) and others do not benefit as much. Also, post-processing of images can be time consuming.
The images below were acquired using the UA instrument. The processing software allows us to create images as widefield, deconvolved (similar in resolution to confocal), and structured illumination. Some of the images below are of single focal planes, while others are of maximum intensity projections created from the entire stack of images.
For more information about this instrument, contact Doug Cromey.
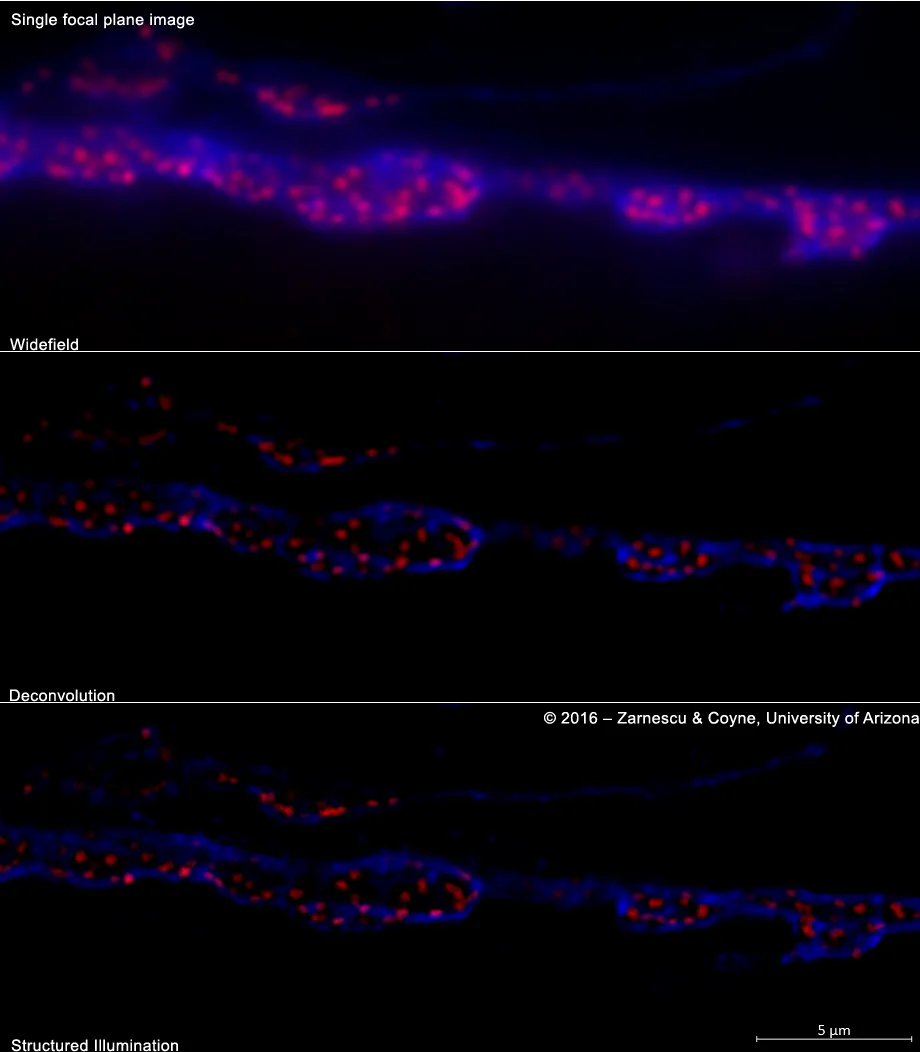
A neuromuscular junction preparation from a third instar Drosophila larva expressing ALS associated mutant TDP-43 in motor neurons immunostained for the neuronal membrane marker hrp (blue) and the active zone protein Bruchpilot (red). - From the laboratory of Daniela Zarnescu, Ph.D., sample prepared by Alyssa Coyne, Ph.D. (Molecular & Cellular Biology, University of Arizona)
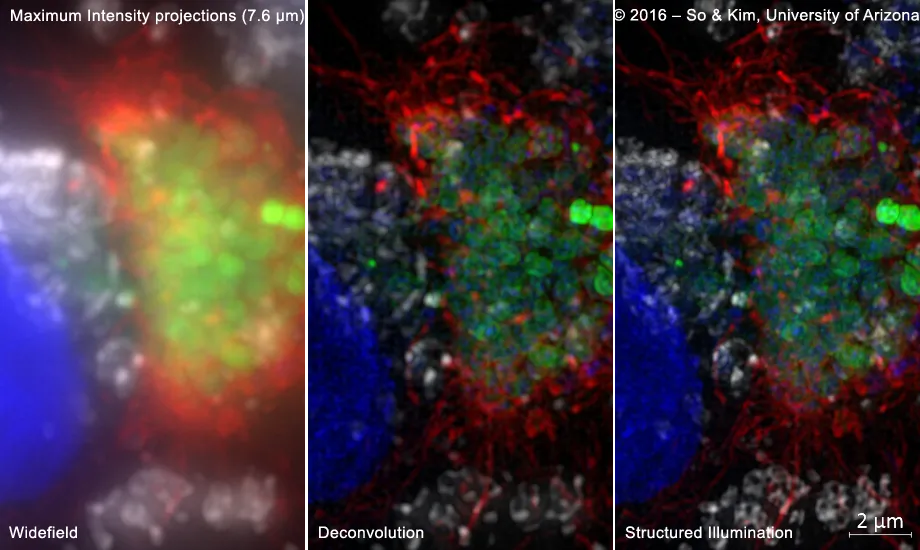
This image shows an epithelial cell infected with human pathogen Neisseria gonorrhoeae. Bacterial cells express GFP (green), and DNA is labeled with DAPI (blue). The long fibers on the bacterial surface (red) are Type IV pili. LAMP1 vesicles in the epithelial cell are in white. - From the laboratory of Magdalene So, Ph.D., sample prepared by Won Kim, B.S. (Immunobiology, University of Arizona)
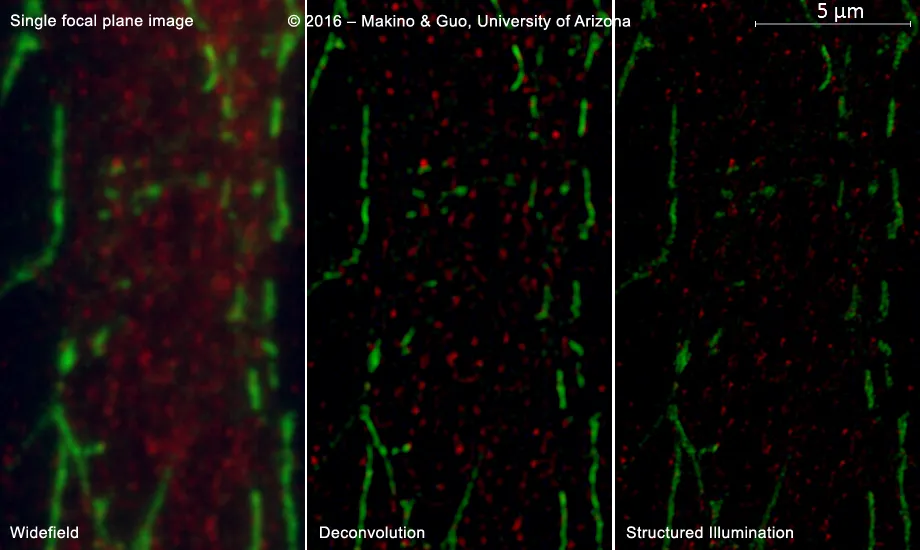
From MPEC cells (mouse pulmonary ECs), with VE-Cadherin (green) and HIF2α (red) labels. - From the laboratory of Ayako Makino, Ph.D., sample prepared by Rui Guo, Ph.D. (Physiology, University of Arizona)
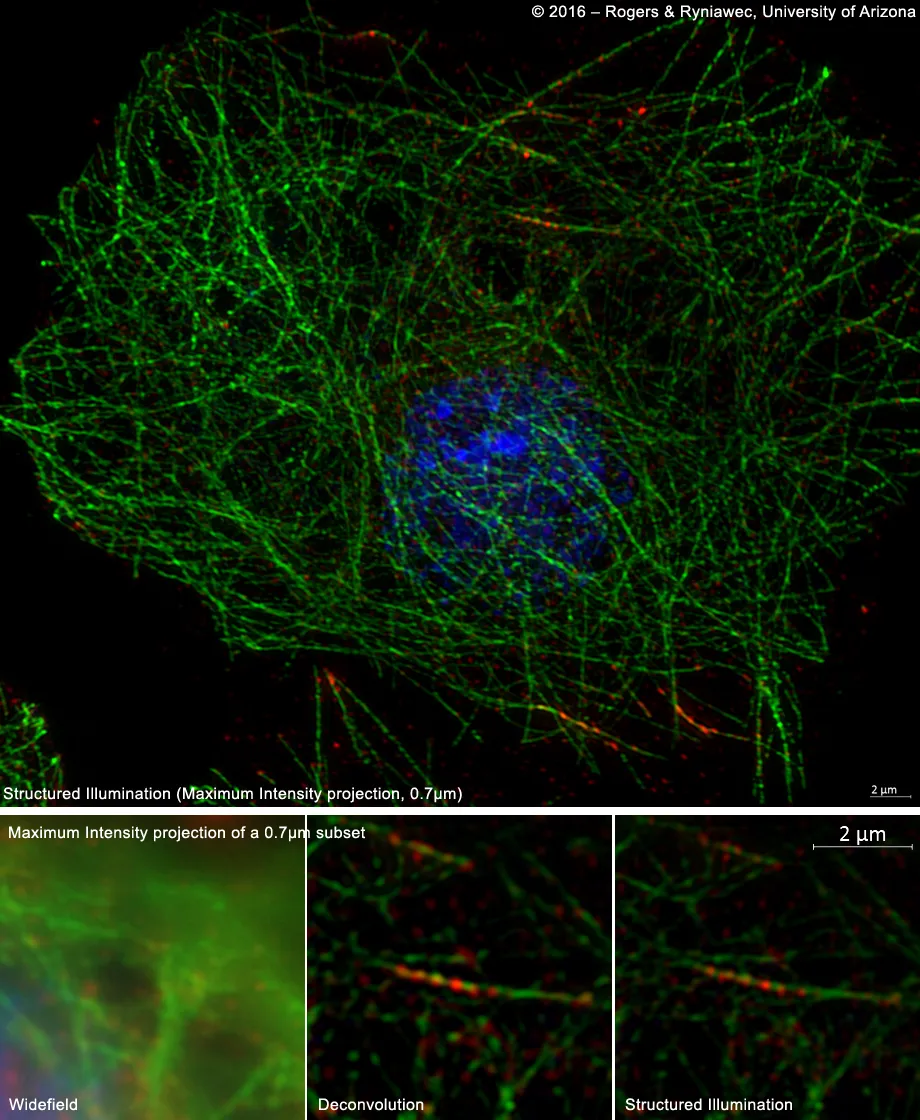
Drosophila Schneider 2 (S2) cell (image size was slightly reduced to fit webpage). The staining is: Blue - Hoescht (DNA), Green - Microtubules (Cy2), and Red - Klp10A (Rhodamine). Klp10A is a kinesin-13 microtubule depolymerase that slides along microtubules and localizes behind +Tips, where it promotes microtubule catastrophe. Inserts show multiple Klp10A molecules localized to a single microtubule. - From the laboratory of Greg Rogers, Ph.D., sample prepared by John Ryniawec, B.S. (Cellualr & Molecular Medicine, University of Arizona)

